Note
Click here to download the full example code
BMI prediction using the COMBO dataset¶
We first consider the COMBO data set and show how to predict Body Mass Index (BMI) from microbial genus abundances and two non-compositional covariates using “filtered_data”.
Import the package¶
import sys, os
from os.path import join, dirname, abspath
classo_dir = dirname(dirname(abspath("__file__")))
sys.path.append(classo_dir)
from classo import classo_problem, clr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
Define how to read csv¶
def csv_to_np(file, begin=1, header=None):
"""Function to read a csv file and to create an ndarray with this
Args:
file (str): Name of csv file
begin (int, optional): First colomn where it should read the matrix
header (None or int, optional): Same parameter as in the function :func:`pandas.read_csv`
Returns:
ndarray : matrix of the csv file
"""
tab1 = pd.read_csv(file, header=header)
return np.array(tab1)[:, begin:]
Load microbiome and covariate data X¶
Load BMI measurements y¶
Normalize/transform data¶
Set up design matrix and zero-sum constraints for 45 genera¶
Set up c-lassso problem¶
Use stability selection with theoretical lambda [Combettes & Müller, 2020b]
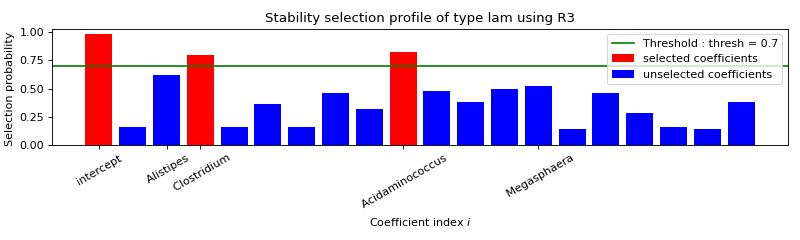
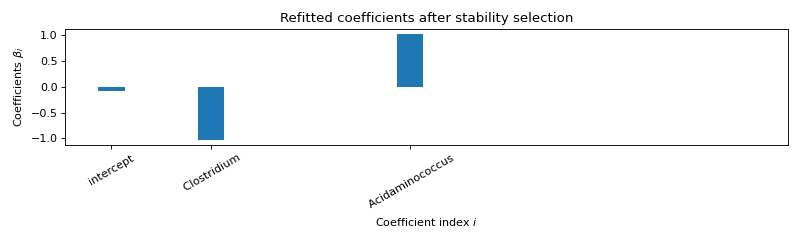
Use formulation R3¶
problem.formulation.concomitant = True
problem.solve()
print(problem)
print(problem.solution)
Out:
FORMULATION: R3
MODEL SELECTION COMPUTED:
Stability selection
STABILITY SELECTION PARAMETERS:
numerical_method : Path-Alg
method : lam
B = 50
q = 10
percent_nS = 0.5
threshold = 0.7
lam = theoretical
theoretical_lam = 0.2818
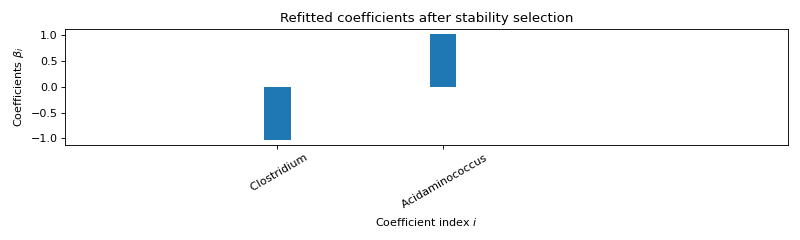
STABILITY SELECTION :
Selected variables : intercept Clostridium Acidaminococcus
Running time : 0.434s
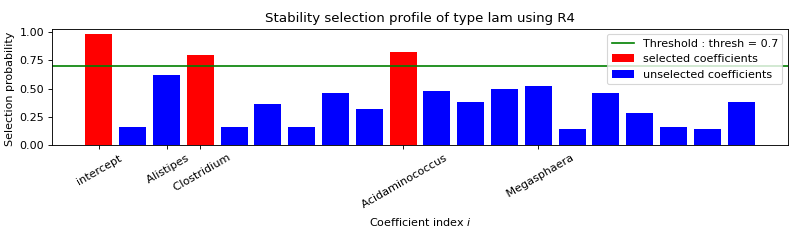
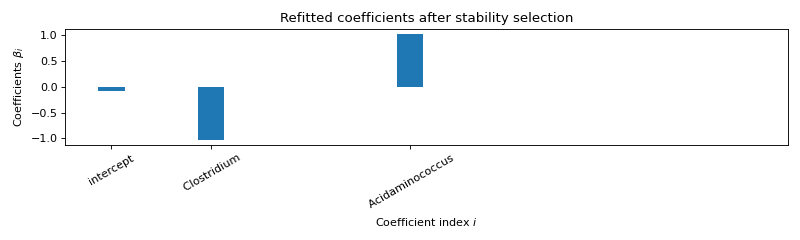
Use formulation R4¶
problem.data.label = label
problem.formulation.huber = True
problem.formulation.concomitant = True
problem.solve()
print(problem)
print(problem.solution)
Out:
FORMULATION: R4
MODEL SELECTION COMPUTED:
Stability selection
STABILITY SELECTION PARAMETERS:
numerical_method : Path-Alg
method : lam
B = 50
q = 10
percent_nS = 0.5
threshold = 0.7
lam = theoretical
theoretical_lam = 0.2818
STABILITY SELECTION :
Selected variables : intercept Clostridium Acidaminococcus
Running time : 0.658s
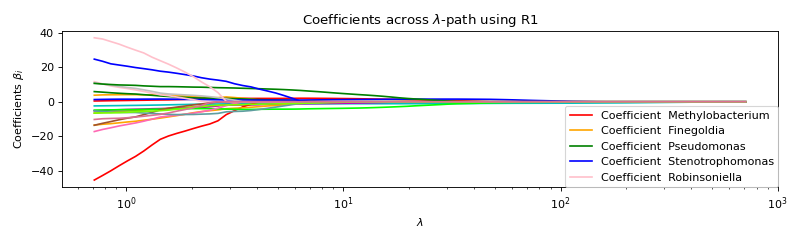
Use formulation R1 with ALO¶
ALO is implemented only for R1 without intercept for now.
problem.data.label = label
problem.formulation.intercept = False
problem.formulation.huber = False
problem.formulation.concomitant = False
problem.model_selection.ALO = True
problem.solve()
print(problem)
print(problem.solution)
Out:
FORMULATION: R1
MODEL SELECTION COMPUTED:
ALO
Stability selection
ALO PARAMETERS:
numerical_method : Path-Alg
lamin = 0.001
Nlam = 80
STABILITY SELECTION PARAMETERS:
numerical_method : Path-Alg
method : lam
B = 50
q = 10
percent_nS = 0.5
threshold = 0.7
lam = theoretical
theoretical_lam = 0.2818
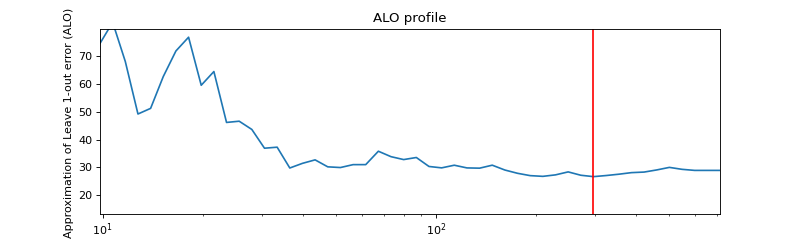
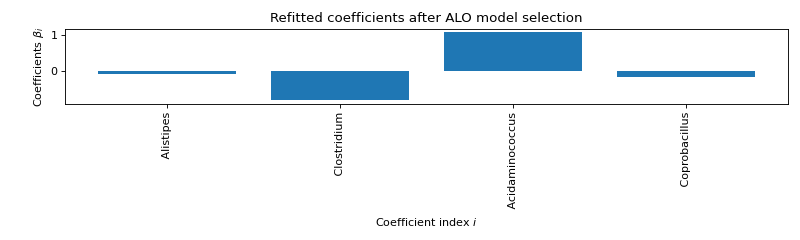
ALO COMPUTATION :
Selected variables : Alistipes Clostridium Acidaminococcus Coprobacillus
Running time : 0.157s
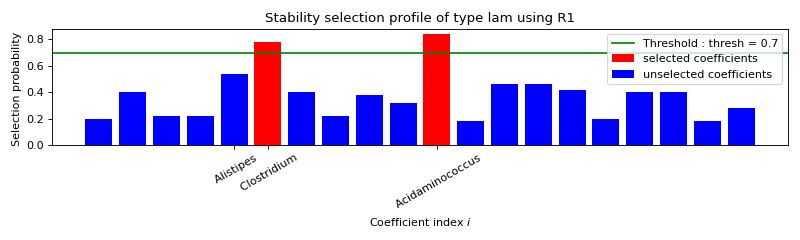
STABILITY SELECTION :
Selected variables : Clostridium Acidaminococcus
Running time : 0.307s
Total running time of the script: ( 0 minutes 3.344 seconds)